Research
Our lab investigates the molecular, cellular, and genetic mechanisms underlying human development and disease by integrating computational biology, organoid technology, and genome topology. By combining large-scale data integration with mechanistic experimental systems, we aim to uncover fundamental regulatory principles and translate them into therapeutic strategies.
Integrating Large-Scale Biological Datasets to Decode Cellular States
We integrate multi-sample and multi-condition datasets to define conserved cellular states and regulatory programs across development and disease.

-
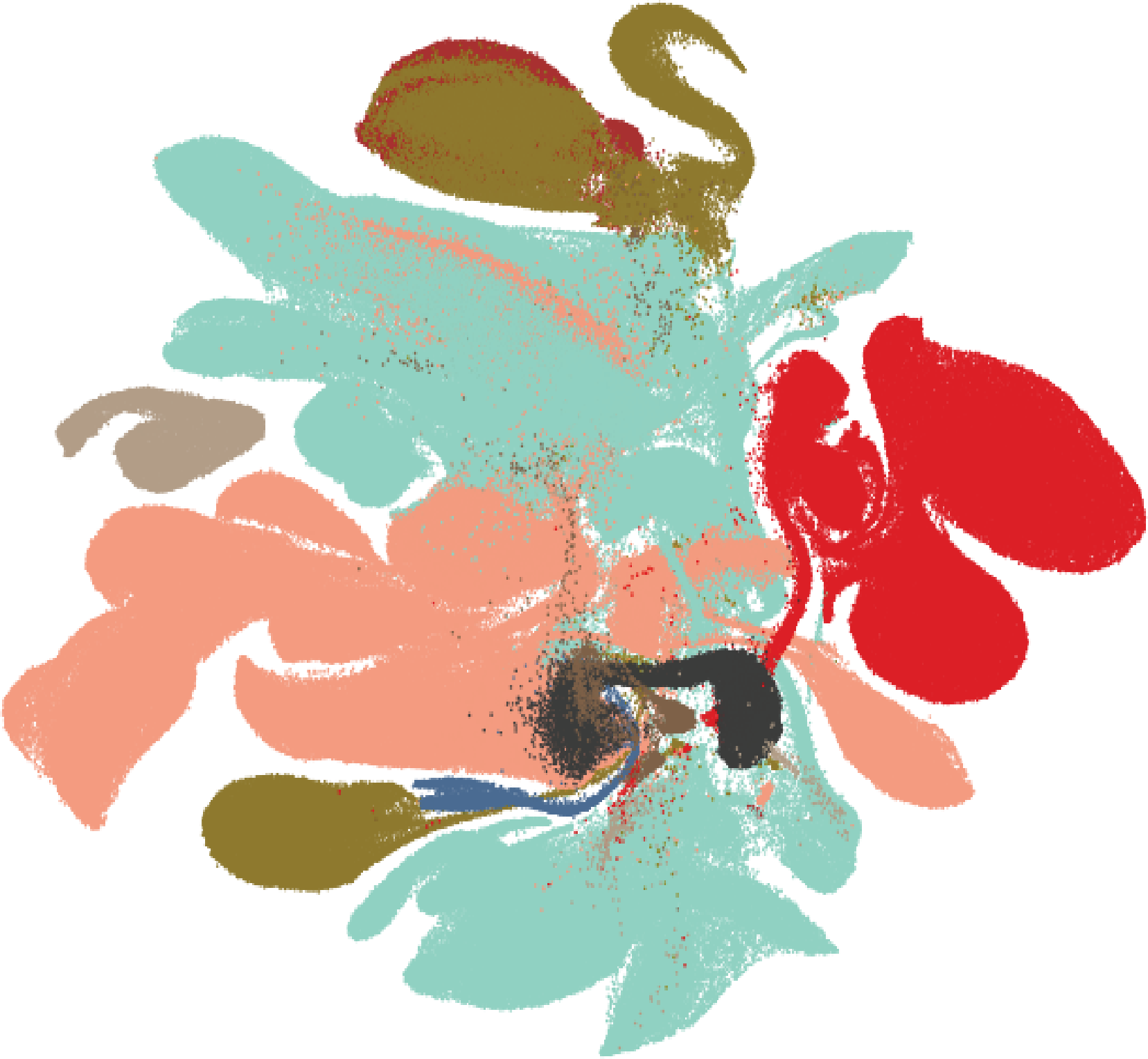
Human Endoderm Organoid Cell Atlas (HEOCA) (Nature Genetics, 2025). We established a single-cell atlas comprising ~1 million cells across diverse endoderm-derived organoid models, including lung, liver, pancreas, intestine, and stomach. HEOCA enables systematic benchmarking of organoid fidelity against primary tissues, reveals ontogenic cell states captured in vitro, and facilitates the identification of disease-associated cellular alterations.
-
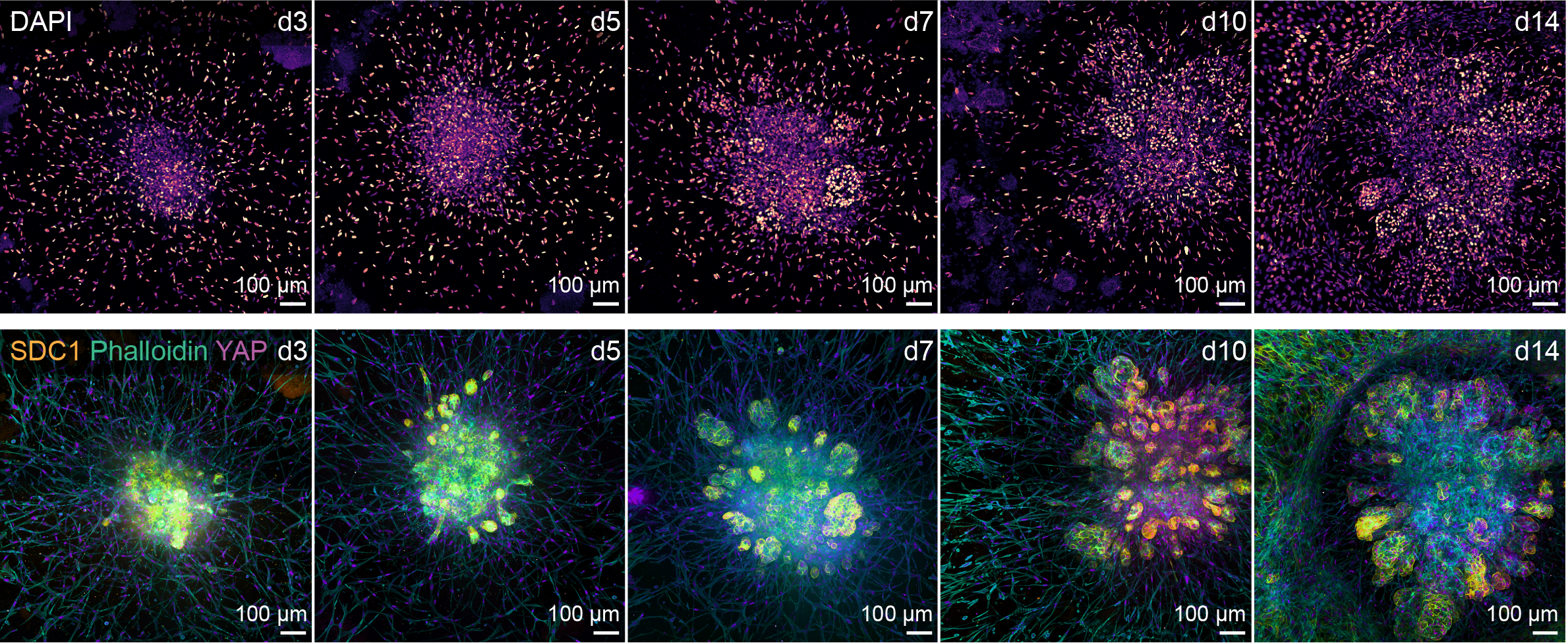
Pancreatic Cancer Atlas (bioRxiv, 2025a). By integrating single-cell transcriptomes from 200 PDAC patient samples, we defined conserved cancer cell and cancer-associated fibroblast (CAF) states and their signaling interactions. Coupling the atlas with patient-derived tumoroids enabled identification of the CAF-responsive receptor Syndecan-1 (SDC1) as a potential therapeutic vulnerability.
Engineering Organoid Models to Recapitulate Human Biology
A central focus of our lab is the design and refinement of organoid systems that faithfully model human tissue architecture and function.
-
Microfold Cell–Enriched Intestinal Organoids (Nature Genetics, 2025). We developed an intestinal organoid system enriched for microfold (M) cells through TNF stimulation, enabling the study of epithelial–immune interactions.
-
Adrenal Cortex Organoids (in revision, 2025). We established a stable human fetal adrenal cortex organoid model to study steroidogenesis and zonation, revealing how WNT, BMP/TGFβ, and cAMP signaling coordinate cortical identity and hormone production.
-
Bioengineered Human Colon Organoids (Cell Stem Cell, 2024). We generated bioengineered colon organoids with in vivo–like cellular complexity and functional architecture.
-
Epidermal Differentiation Models (PNAS, 2019). Using iPSCs from EEC syndrome patients carrying TP63 mutations, we uncovered aberrant mesodermal activation that impaired epidermal commitment and demonstrated partial rescue through pathway inhibition.
From Organoids to Therapeutic Insight
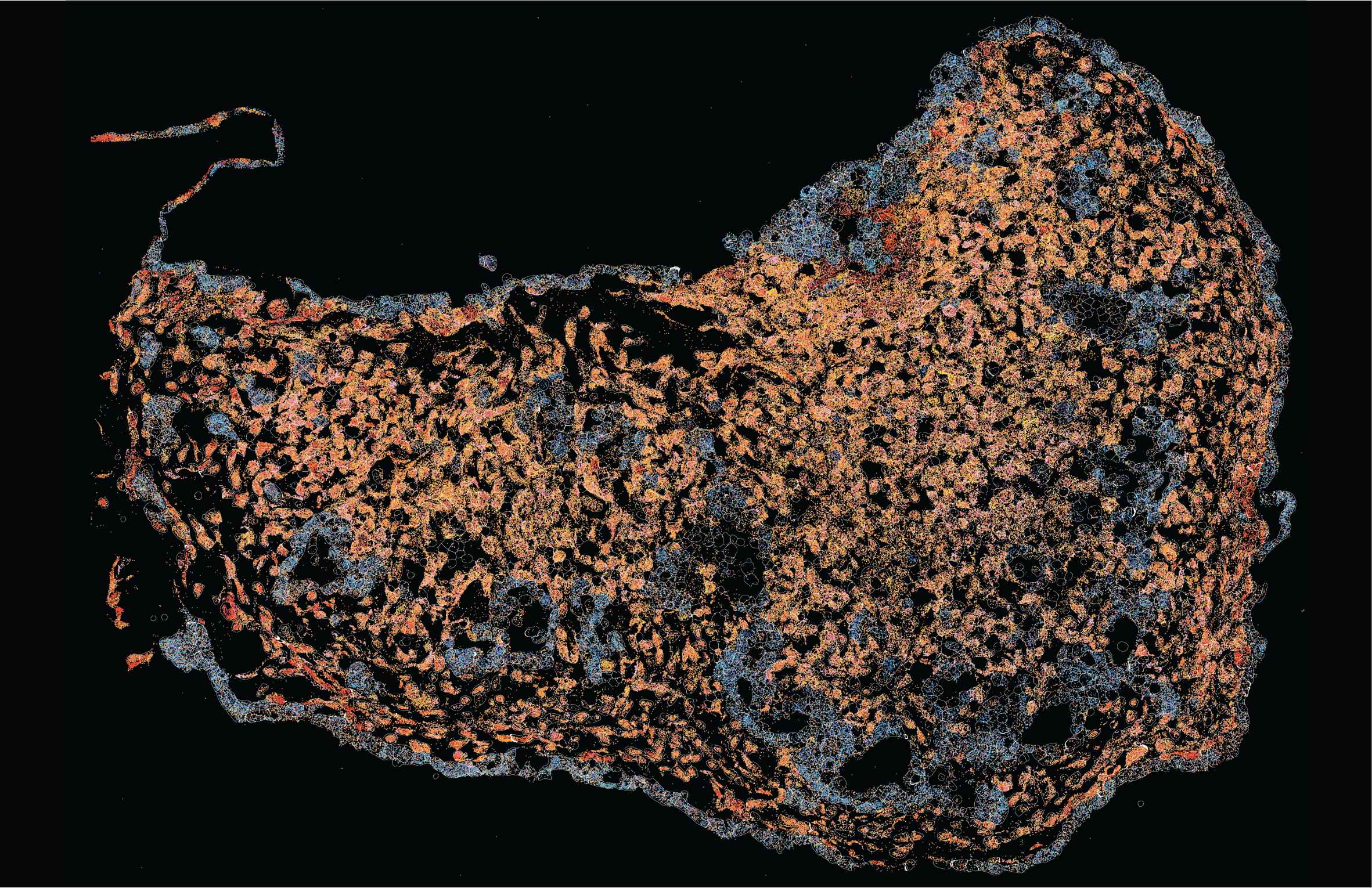
We use organoid systems not only to model human biology, but also to identify actionable mechanisms and therapeutic strategies.
-
Pancreatic Tumoroids (bioRxiv, 2025a, 2025b). Our modular patient-derived tumoroids reconstruct tumor–stroma interactions and ductal architecture, providing a platform to investigate immune evasion and evaluate therapeutic combinations.
-
Retina Organoids for Base Editing (Nature Medicine, 2025). We demonstrated high-efficiency base editing in primate and human retinal tissues, highlighting the translational potential of precision genome editing in organoid systems.
Together, these approaches provide a scalable framework linking cellular states, regulatory mechanisms, and therapeutic opportunity.
Developing Computational Frameworks for Regulatory Biology

We develop computational tools to identify regulatory drivers of cell fate determination.
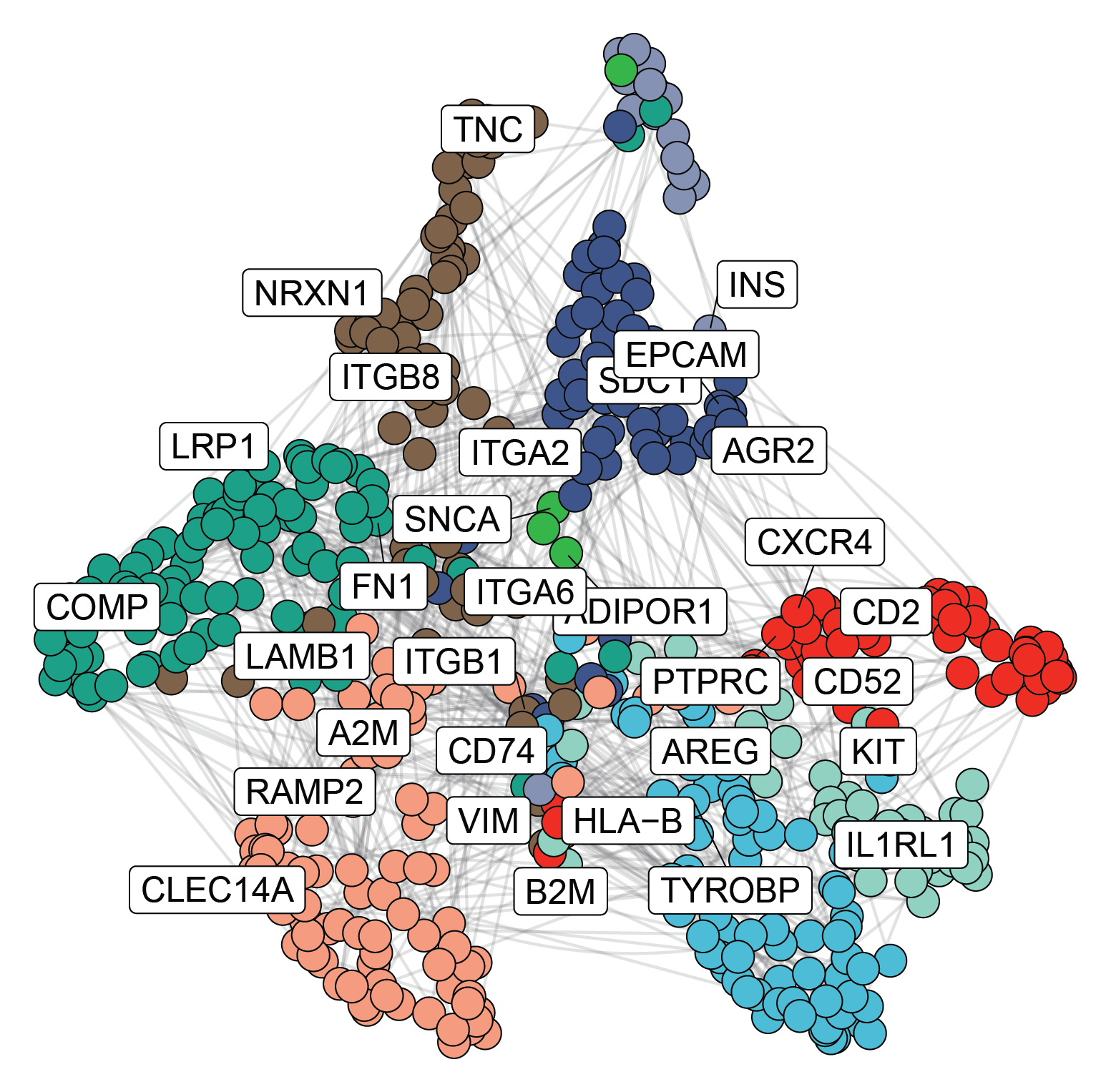
ANANSE (Nucleic Acids Research, 2019) is our enhancer-centered framework for inferring gene regulatory networks (GRNs) and prioritizing key transcription factors (TFs) controlling cell identity. Because cell type–specific TFs predominantly act through enhancers, ANANSE integrates enhancer activity with TF binding motifs to predict genome-wide TF binding profiles and construct cell type–specific regulatory networks. By comparing differential networks between cell states, ANANSE identifies candidate TFs driving cell fate transitions and consistently outperforms promoter-based approaches in predicting factors sufficient for trans-differentiation. Using this framework, we generated a cross-tissue atlas of key transcription factors across 18 human tissues. ANANSE is open source and widely applicable for studying transcriptional regulation in development and disease.
Decoding 3D Genome Architecture

We investigate how genome topology shapes gene regulation (Cell, 2015). CTCF and the cohesin complex are central regulators of higher-order chromatin organization in mammalian genomes. Using CRISPR/Cas9-mediated genomic editing combined with chromosome conformation capture approaches, we demonstrated that the location and relative orientation of CTCF-binding sites (CBSs) determine the specificity of long-range chromatin looping. By inverting CBS elements at endogenous loci, including the protocadherin and β-globin regions, we showed that CTCF site orientation reconfigures enhancer–promoter interactions, alters chromatin domain architecture, and reshapes gene expression patterns. These findings provide functional evidence that three-dimensional genome organization is encoded, in part, by the linear orientation of regulatory elements, offering mechanistic support for the loop extrusion model of chromatin folding.